The first tutorial is supplied for carrying out the full pathway of the ab-initio crystal structure solution process, from indexing up to the structure solution by Direct Methods, in the case of cimetidine.
The only required information by EXPO is the experimental powder diffraction pattern and the chemical formula C10H16N6S.
The examples directory contains the necessary files:
- cime.exp: input file for EXPO
- cime.dat: synchrotron X-ray diffraction data file containing the initial 2-theta, the 2-theta step and the final 2-theta values (in the first line) and the experimental diffraction counts (in the rest of lines).
To access to examples directory:
- Open EXPO by double clicking on Expo2 icon.
- Click on File > Load Examples
and select the file cime.exp.
The content of the file cime.exp follows.
%structure cime %job cimetidine (C10H16N6S) -- Synchrotron data %data pattern cime.pow wavelength 1.52904 synchrotron %ntreor %continue
It contains the command (%data) for providing the minimal required information (name of data file, wavelength and type of radiation used to collect data, if different from laboratory X-rays) and the commands for activating the indexing process (%ntreor) and the ab-initio structure solution (%continue). This file can be visualized and eventually modified by the ‘Edit’ button.
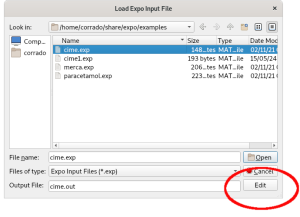
More information about the creation of input file can be found in the section Preparing input file.
Clicking on OK, the powder diffraction data is displayed.
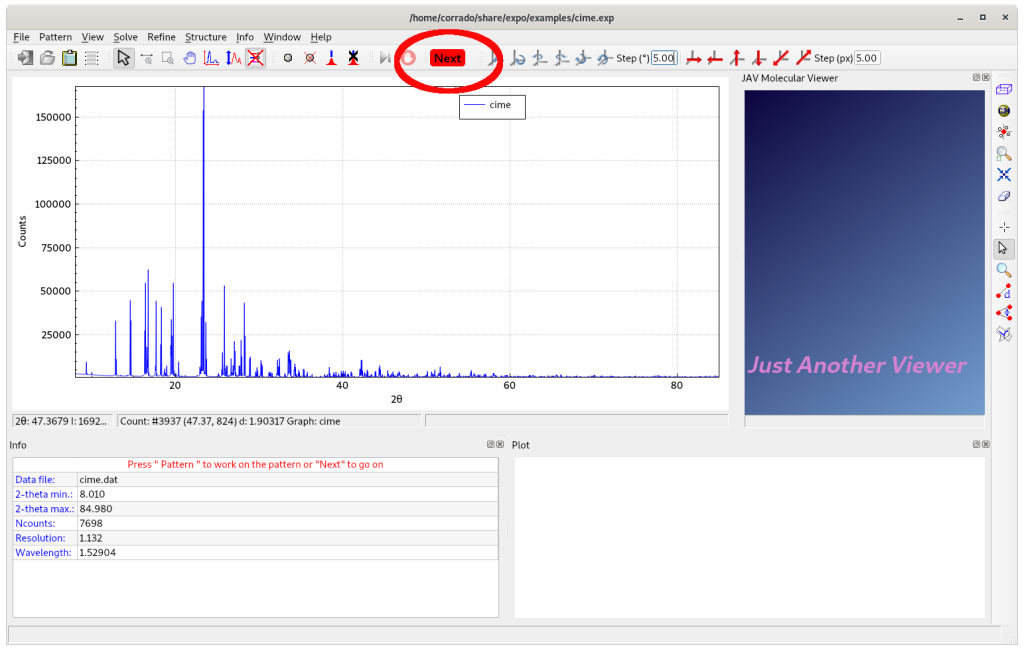
Selecting Next, an automatic peak search procedure is executed. The d-values corresponding to the detected experimental peaks are automatically supplied to the indexing procedure. At the end of this process, EXPO provides the indexing results by graphic interface:
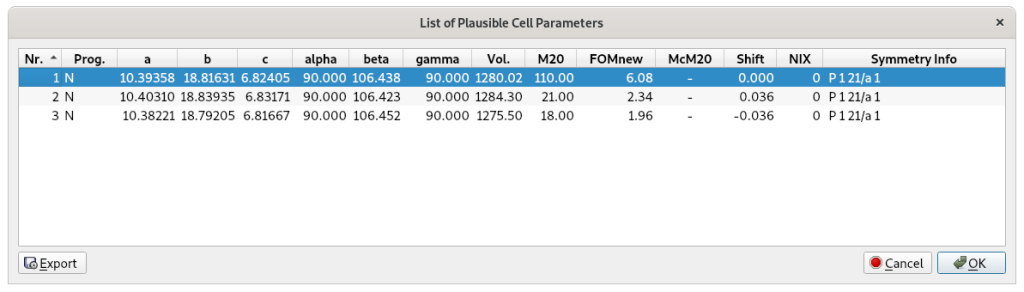
Click on OK to accept the selected plausible cell parameters and to move to the next step: the space group determination.
The space group determination step, as well as direct methods, requires the unit cell content information. It can be easily derived by taking into account the structure formula and the cell volume (Vcell) which is evaluated, by EXPO, from the cell parameters. (The cell volume is graphically provided and/or contained in the output file of EXPO).
The cell content is obtained by multiplying Z by the structure formula. You can calculate Z as the ratio between Vcell and the molecular volume (Vmol): Z = Vcell/Vmol.
Vmol can be calculated by one of the two following alternative ways:
- Using the 18 Å3 rule: counting all the non-hydrogen atoms in the chemical formula and multiplying this number by 18 Å3;
- Using the following approximate atomic volumes: 14 Å3 for C, 12 Å3 for N, 11 Å3 for O, 5 Å3 for H, 26 Å3 for Cl, 25 Å3 for S.
In case of cimetidine compound, the chemical formula is C10H16N6S and the cell volume is 1280 Å3. If the first criterion is applied, the molecular volume is given by: Vmol = 18 x 17 = 306 Å3. Using the second approach Vmol = 14 x 10 + 5 x 16 + 12 x 6 + 26 x 1 = 318 Å3.
The corresponding Z value is: Z = Vcell/Vmol = 1280/306 = 4.18, that can be approximated to 4. So 4 molecules of Cimetidine are in the unit cell and the unit cell content to be supplied to EXPO by graphic interface is (C10H16N6S)4 or C40H64N24S4.
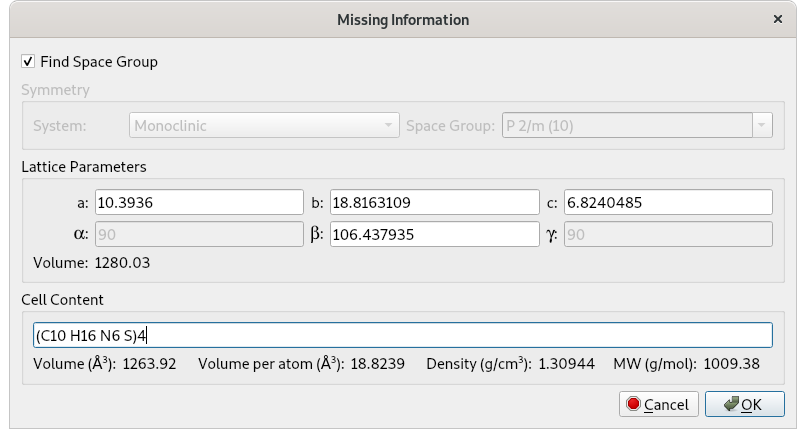
Click on OK in order to perform the space group determination and then on the Next button to go on continuously. At the end of the procedure, a list of the space groups and extinction groups, compatible with the lattice symmetry and ranked according probability values, is given:
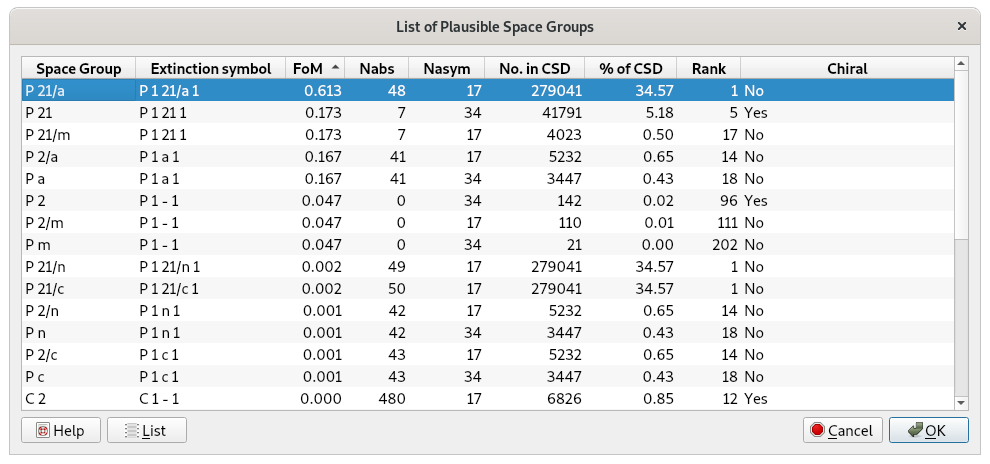
The most plausible extinction group P 1 21/a 1 is automatically selected and by clicking on OK, the space group P21/a, corresponding to the chosen extinction group, is considered.
After the determination of the space group, EXPO passes to the extraction step, which is carried out by using the selected space group. Click on the Next button continuously. At the end of the process, a list of integrated intensity values, extracted from the experimental powder pattern, is obtained and automatically supplied to the Direct Methods process for structure solution.
The final result is visualized on JAV Molecular Viewer.
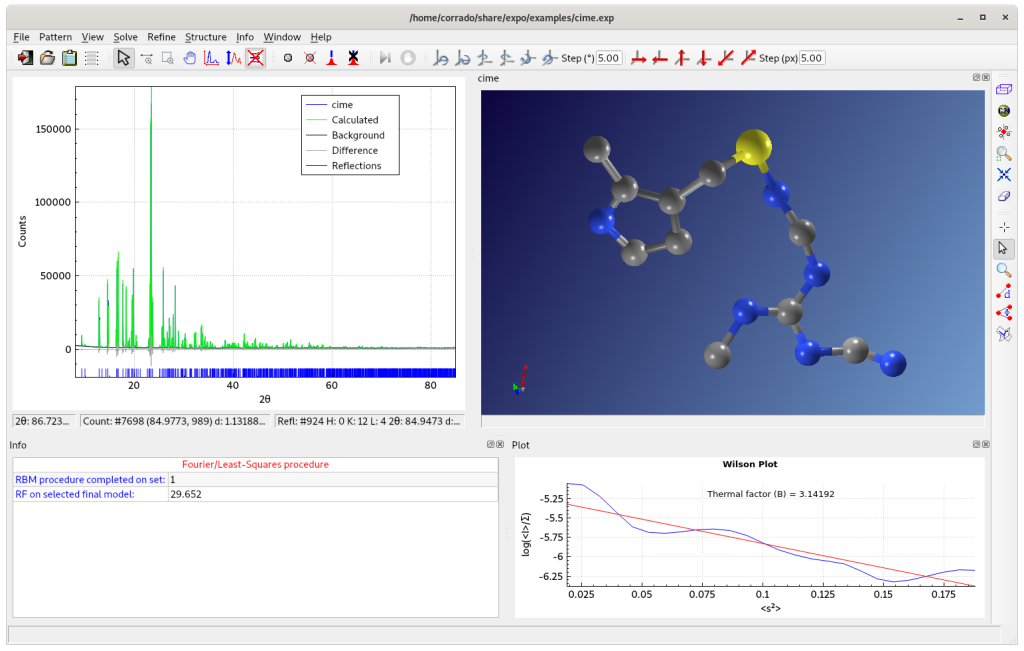
Some errors in the assignment of carbon or nitrogen atomic species can be easily recognized and corrected by using the graphic option ‘Modify’ from JAV menu.
The aim of this example is to guide the user in case of failure of a default run of EXPO.
Open EXPO by double click on Expo icon.
Click on File > Load Examples
Select the input file merca.exp.
%structure merca %job 2-Mercaptobenzoic acid %data cell 7.885 5.976 14.949 90.0 100.48 90 spacegroup p 21/c content (C 7 O 2 S 1 H 6)4 pattern merca.xy wavelength 1.54056 %continue
In this example, it is assumed that the cell and space group have been already determined and the unit cell content corresponds to 4 molecule of C7H6O2S.
Click on the Next button to go on continuously.
The final structure model provided by EXPO could not correspond to the correct crystal structure of 2-Mercaptobenzoic acid (the model is not chemically interpreted). This can occur when the most plausible set of phases, provided by Direct Methods, is unreliable.
In a default run of EXPO, Direct Methods generate several phasing trials; not more than 20 of them are stored and ranked according to decreasing values of a suitable combined figure of merit (CFOM). The largest CFOM set of phases is automatically selected for structure solution. In case of failure, the procedure ALLTRIALS can be applied: all the rest of stored phasing trials are explored in order to find the correct solution.
This can be automatically performed by graphic interface:
a) Selecting from the top-level menu Solve and then Explore Trials
b) Clicking on the Select all new trials button
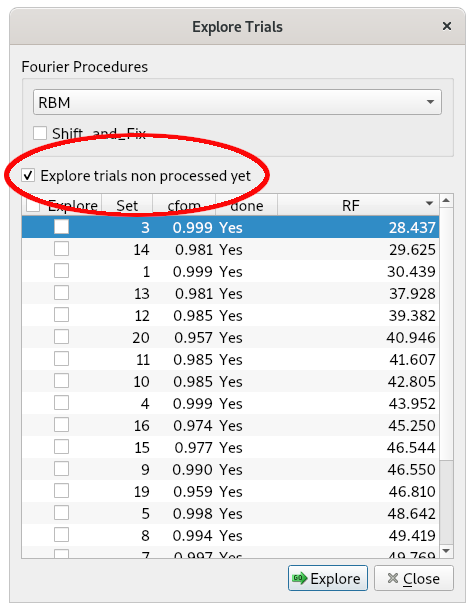
c) Clicking on GO
At the end of the procedure, EXPO ranks the phasing trials (and the corresponding structure models) according to the increasing RF agreement factor value.
The best solution (corresponding to the lowest RF value) is selected and displayed.
The purpose of this tutorial is to guide the user through the crystal structure solution process by Direct Space methods of paracetamol (form I polymorph) compound.
Open EXPO, select Load Examples.
Select the input file paracetamol.exp.
Clicking on the Next button, the file paracetamol.mol, containing a 3D molecular description of paracetamol, is loaded and displayed and the following window appears:
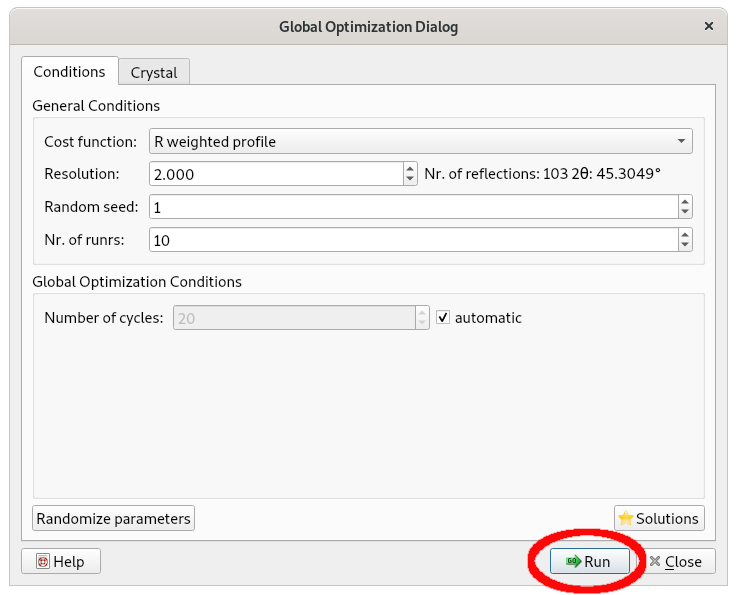
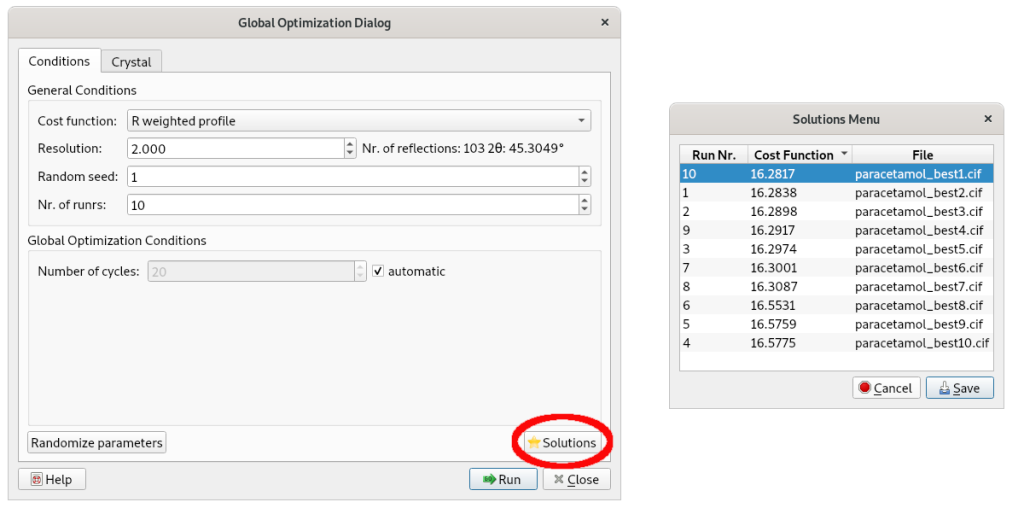
Select Run; the Simulated Annealing procedure is applied performing 10 runs whose corresponding final structure models can be explored by the button Solutions.
Cimetidine – Cernik, R.J., Cheetham, A.K., Prout, C.K., Watkin, D.J., Wilkinson, A.P., Willis, B.T.M. (1991). J. Appl. Cryst. 24, 222-226.
2-Mercaptobenzoic acid – Steiner, T. (2000). Acta Cryst. C56, 876–877.
Paracetamol (form I polymorph) – Nichols, C. & Frampton, C. S. (1998). J. Pharm. Sci.87, 684–693.